Description
These tracks contain the results of DNase I hypersensitivity experiments performed by the
John Stamatoyannapoulos lab
at the University of Washington from September 2007 to January 2011, as part of the
ENCODE project first production phase.
Colors were assigned to cell types based on similarity of signal.
Other views of this data (along with additional documentation) are available from the hg19
ENCODE UW DNaseI HS track.
Display Conventions and Configuration
This track is a composite annotation track containing multiple subtracks, one for each cell type.
The display mode and filtering of each subtrack can be individually controlled.
For more information about track configuration, see
Configuring Multi-View Tracks.
Methods
Raw sequence data files were processed by the UCSC ENCODE DNase analysis pipeline (July 2014 specification), diagrammed here:
 Credit: Qian Alvin Qin, X. Liu lab
Credit: Qian Alvin Qin, X. Liu lab
Briefly, sequence files were aligned to the hg38 (GRCh38) genome assembly augmented with 'sponge'
sequence (ref). Multi-mapped reads were removed, as were reads that aligned to 'sponge' or
mitochondiral sequence. Results from all replicates were pooled, and further processed by
the Hotspot program to call peaks as well as broader regions of activity ('hotspots'), and to
create signal density graphs.
Signal graphs were normalized so the average value genome-wide is 1.
The cell types were clustered into a binary tree, a rainbow was cast to the leaf nodes providing coloring based on similarity.
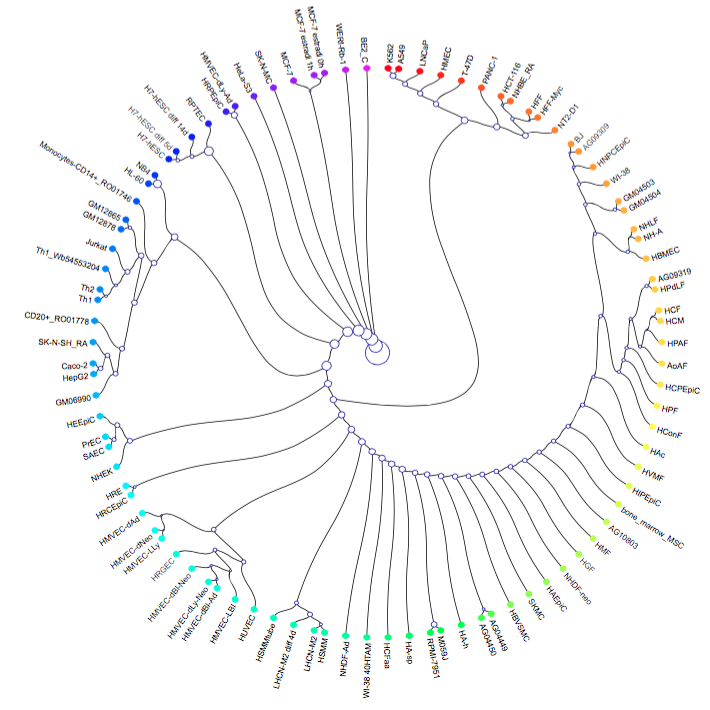 Credit: Chris Eisenhart, J. Kent lab
Credit: Chris Eisenhart, J. Kent lab
(Please note there is different coloring on the ENCODE hg38
Transcription track,
Layered H3K4Me1 track,
Layered H3K4Me3 track, and
Layered H3K27Ac track,
which match the coloring used in their previous versions lifted from the hg19 assembly).
Credits
The processed data for this track were produced by UCSC. Credits for the primary data
underlying this track are included in the
ENCODE UW DNaseI HS track
description.
References
Miga KH, Eisenhart C, Kent WJ.
Utilizing mapping targets of sequences underrepresented in the reference assembly to reduce false
positive alignments.
Nucleic Acids Res. 2015 Nov 16;43(20):e133.
PMID: 26163063
Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, Sheffield NC, Stergachis AB, Wang
H, Vernot B et al.
The accessible chromatin landscape of the human genome.
Nature. 2012 Sep 6;489(7414):75-82.
PMID: 22955617; PMC: PMC3721348
See also the references in the
ENCODE UW DNaseI HS
track.
 US Server
US Server European Server
European Server Asian Server
Asian Server